Ciliates are a large group of unicellular eukaryotes that have a highly unusual genome architecture. They have two, a micronucleus (MIC) and a macronucleus (MAC). The MIC is where genes are stored and it is passed on during sexual reproduction but it does not produce mRNA. The MAC is not passed during sexual reproduction, it is much larger than the MIC containing several copies of each gene and it is derived from the MIC (1). Ciliates normally divide asexually and in that case both nuclei are duplicated and one copy is passed on to each daughter cell. The reason for this genome architecture seems to be the very large cell size of Ciliates making it necessary to produce large amounts of proteins from each gene; this makes it necessary to have several copies of each gene from which mRNA can be produced.
(Click for bigger)
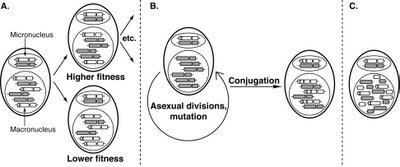
“Genome architecture drives protein evolution in ciliates through the impact of selection operating on processed chromosomes in a somatic nucleus that divides by amitosis. Each ciliate contains a germline micronucleus, with a canonical eukaryotic genome, and a somatic macronucleus, represented by a large polyploid nucleus. A. If a deleterious mutation occurs (shown as an X), the chromosome carrying that mutation can be lost following unequal assortment during amitosis of the macronucleus. While this mutation may eventually be completely eliminated from the macronucleus, it will be present in the micronucleus. B. During subsequent rounds of asexual division, the micronucleus will acquire additional mutations. Given sufficient time and/or population size, one or more of these mutations may be compensatory. After conjugation, individuals with compensatory mutations can increase in frequency in the population. C. These processes are exaggerated in ciliates with extensively fragmented genomes, where every allele and locus is able to assort independently.”
Its and interesting theory although a lot more data will have to be accumulated before it is validated or another explanation can be found for the rate of evolution in Ciliates.
Although ‘Developmentally Regulated Genome Rearrangements’ seem very odd, they are fairly common (3). For example Genome-wide rearrangements has been found in Nematodes, Copepods, Hagfish Foraminifera and Ciliates and Targeted rearrangements are found in places like the vertebrate immune system. It is likely that this phenomenon does have an effect on the rate of evolution and it seems that this area of study will reveal interesting results.
Refs:
GENOME REMODELING IN CILIATED PROTOZOA
Annual Review of Microbiology Vol. 56: 489-520
doi:10.1146/annurev.micro.56.012302.160916
2) Rebecca A. Zufall, Casey L. McGrath, Spencer V. Muse, and Laura A. Katz
Genome Architecture Drives Protein Evolution in Ciliates
MBE Advance Access published on
http://mbe.oxfordjournals.org/cgi/content/abstract/msl032v1
Evolution of Developmentally Regulated Genome Rearrangements in Eukaryotes
JOURNAL OF EXPERIMENTAL ZOOLOGY (MOL DEV EVOL) 304B (2005)
http://www3.interscience.wiley.com/cgi-bin/abstract/110572604/ABSTRACT
1 comment:
Do you like playing in the game which you need to use wow gold, when you do not have World of Warcraft Gold, you must borrow warcraft gold from friends, or you buy wow gold. If you get cheap wow gold, you can continue this game.
Post a Comment